Collaborators
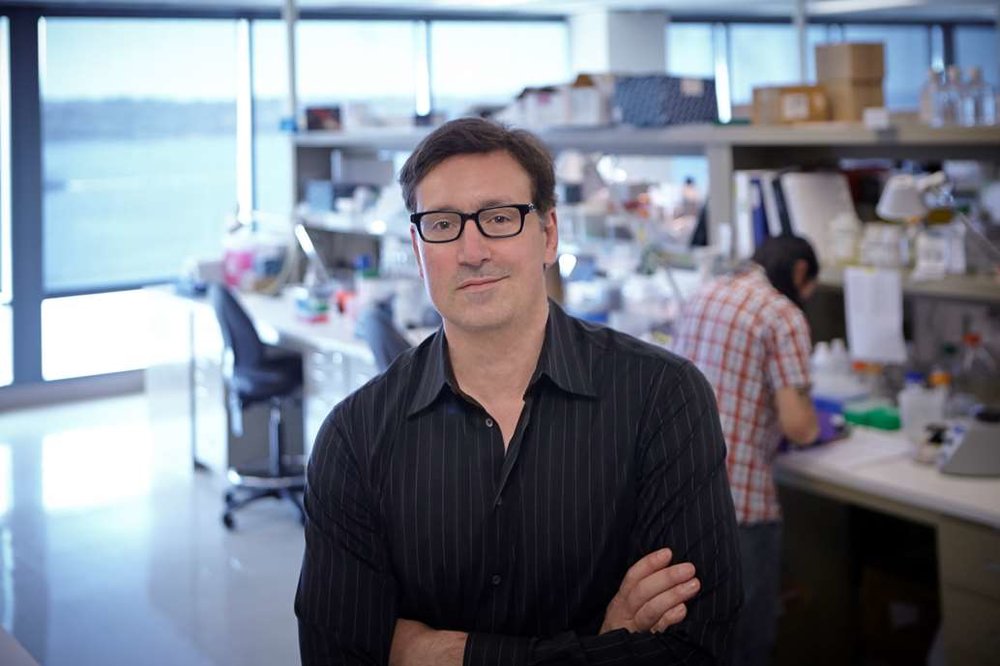

Dr. Stamatoyannopoulos' laboratory uses high-scale molecular, computational, and genome engineering technologies to decode the regulatory circuitry of the human and other complex genomes. Major ongoing efforts are (i) to create comprehensive atlases of regulatory DNA encoded in the human and mouse genomes; (ii) to define regulatory networks that control cell fate and response; (iii) to identify and characterize human regulatory variants associated with common human diseases; and (iv) to develop next-generation technologies for interrogating, interpreting, and programming the human regulatory genome.
Dr. Stamatoyannopoulos holds degrees in Biological Sciences, Symbolic Systems, and Classics from Stanford University, and an M.D. from the University of Washington School of Medicine. Dr. Stamatoyannopoulos completed his medical training at Harvard Medical School, including internship and residency in Internal Medicine at Brigham and Women's Hospital, and fellowship in Oncology and Hematology at the Dana Farber Cancer Institute and Massachusetts General Hospital.

Following a few semesters studying philosophy and history at Freiburg University and Munich University, Cremer studied physics in Munich (with financial support from the Studienstiftung des Deutschen Volkes) and completed his Ph.D. in genetics and biophysics in Freiburg. This was followed by post-doctoral studies at the Institute for Human Genetics at Freiburg University, several years in the United States at the University of California, and his "Habilitation" in general human genetics and experimental cytogenetics at Freiburg University. Since 1983, he is teaching as a professor (chair since 2004) for "applied optics and information processing" at the Kirchhoff Institute for Physics at the University of Heidelberg. In addition, he is a member of the Interdisciplinary Center for Scientific Computing of the Institute for Pharmacy and Molecular Biotechnology, as well as of the University’s "Bioquant" Center. Cremer is a participant in three current "Projects of Excellence" of the University of Heidelberg (2007–2012), and is also a partner in the Biotechnology Cluster for cell-based and molecular medicine, one of five clusters selected in 2008 as German BMBF Clusters of Excellence. Elected as Second Speaker of the Senate of the University of Heidelberg (since 2006), Cremer is also involved in university governance and politics. In his function as adjunct professor at the University of Maine and as member of the Jackson Laboratory (Bar Harbor, Maine), where he undertakes research for several weeks each year during the semester breaks, he is involved in the establishment of the biophysics center (Institute for Molecular Biophysics), which is linked with the University of Heidelberg through a "Global Network" collaboration.

Thomas Cremer is Professor of Human Genetics and Anthropology within the Ludwig-Maximilians-University (LMU) in Munich, Germany. Research within the Cremer laboratory is focused on Molecular Cytogenetics and 3D/4D analyses of nuclear structure using confocal microscopy and live cell imaging. Researchers also use three-dimensional Fluorescence in Situ Hybridization (3D FISH) to study nuclear architecture.
Thomas studied medicine at the Albert-Ludwigs-University of Freiburg. Along with his physicist brother, Christoph Cremer, he showed an early interest in nuclear architecture and cutting edge microscopy. Together they pioneered laser-UV-micro-irradiation experiments that indirectly implied a territorial organization of chromosomes in the interphase nucleus. Thomas has been chair of the department of Human Genetics and Anthropology at the University of Munich since 1995 and is a member of the German Academy of Sciences Leopoldina (2006).
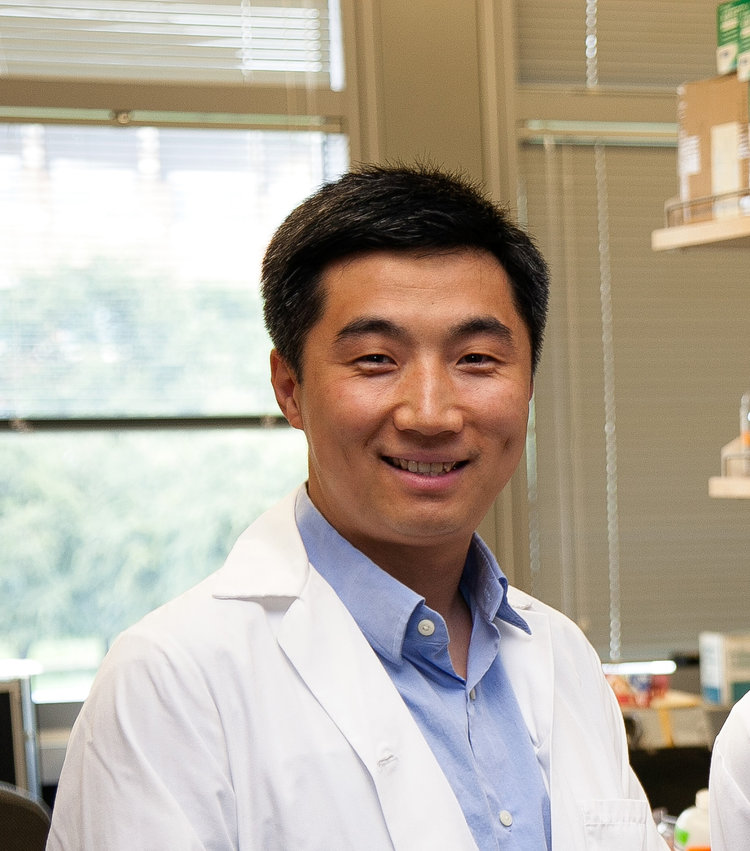

The Gao research group is interested in molecular engineering and nanomedicine, using engineered nanostructures for detection, analysis and treatment of human diseases such as cancer, cardiovascular diseases, infectious diseases and neurological diseases.
Nanotechnology has recently received broad interest from biomedical researchers and may change the very foundations of disease diagnosis, treatment and prevention. From technology development side, currently we are developing semiconductor quantum dots (Qdots) for optical imaging, magnetic nanoparticles for MRI and magnetic manipulation, metallic nanoparticles for photoacoustic imaging and image-guided therapy, and polymeric and natural protein nanoparticles for drug delivery. From the application side, we are interested in molecular screening, diagnosis, prognosis, and treatment. In general, we are interested in any innovative nanotechnology that has impact on biology and medicine.

The goal of the Raj Lab is to develop a quantitative understanding of molecular underpinnings of cellular behavior and then use this knowledge to better human health. Our research combines new tools we have developed for single cell imaging with new genomic and computational methods to make highly accurate measurements of the molecular components underlying cellular function, focusing in particular on the behavior of single cells within an ensemble. Many of the measurement tools that we have developed also have applications in diagnostics for infectious disease and cancer. Also, our results also provides a quantitative basis for synthetic biology. Areas of particular interest include cancer biology, long non-coding RNA, quantitative genetics, stem cell biology with applications to cartilage regeneration and diagnostics based on ultra-rapid viral detection.

Dr. Akilesh is presently applying next generation genomic tools and analyses to understand the development, structure and function of the kidney and its component cells in health and disease. He has received the Damon Runyon Cancer Research Fellowship (2013-2016) to support these studies.

Dr. Hoofnagle’s laboratory focuses on using proteomic and metabolomic approaches to investigate the intersection of inflammation and lipid metabolism. His laboratory has built robust assays for the quantification of proteins in lipoproteins using targeted proteomic approaches to investigate the changes in the HDL proteome due to diabetes and obesity. They are pioneering the use of immunoaffinity peptide enrichment strategies of analyte-specific peptides from tryptic digests in the clinical analysis of low-abundance serum proteins. The laboratory has developed assays for small molecule analytes, including arginine, arginine metabolites, and vitamin D, which are being used in basic science and large-scale clinical studies. Several mouse models are being used to assess the importance of complement regulatory proteins in insulin sensitivity and lipid metabolism. All of these studies are in collaboration with outstanding investigators from the University of Washington and elsewhere.

Mike Hawrylycz joined the Allen Institute in 2003. He is responsible for the direction of the data analysis and annotation effort. Hawrylycz has worked in a variety of applied mathematics and computer science areas, addressing challenges in consumer and investment finance, electrical engineering and image processing, and computational biology and genomics. Hawrylycz received his Ph.D. in applied mathematics at the Massachusetts Institute of Technology. He subsequently was a post-doctoral researcher in the Computer Research and Applications Group at the Los Alamos National Laboratory.
Peng joined the Allen Institute in 2012 to build a computational neuroanatomy and smart imaging group for the Institute's new initiatives in neural coding and cell types. His current research focuses on bioimage analysis, large-scale informatics, machine learning, as well as computational biology. Before joining the Allen Institute, Peng was the head of a computational bioimage analysis lab at Howard Hughes Medical Institute, Janelia Farm Research Campus. His recent work includes developing novel algorithms for 3-D+ image analysis and data mining, building single-neuron whole-brain level 3-D digital atlases for model animals, and Vaa3D (http://vaa3d.org), which is a high-performance visualization-assisted analysis system for large 3-D+ biological and biomedical-image datasets. He is also the inventor of the widely cited minimum-redundant maximum-relevance (mRMR) feature selection algorithm in machine learning. Peng received his Ph.D. in biomedical engineering from Southeast University, China. He held postdoctoral positions at the Lawrence Berkeley National Laboratory at the University of California, Berkeley (computational biology, bioinformatics, and high-performance data mining with a particular focus on gene expression analysis) and at Johns Hopkins University Medical School (human brain imaging and analysis). He won several awards, including a Cozzarelli Prize (2013) for his collaborative research on dragonfly neurons, which "recognizes outstanding contributions to the scientific disciplines represented by the National Academy of Sciences (USA)".